Asthma may heighten flu risk and cause dangerous mutations
A subtype of asthma in adults may cause higher susceptibility to influenza and could result in dangerous flu mutations.
University of Queensland-led animal studies have found that paucigranulocytic asthma (PGA) – a non-allergic form of the condition – allows the flu virus to flourish in greater numbers in sufferers.
UQ PhD candidate Katina Hulme said this was due to the asthma’s suppression of the immune system.
“We were first tipped off about this during the 2009 swine flu pandemic,” Ms Hulme said.
“Asthma was identified as the most common underlying medical condition in individuals hospitalised with flu, and these individuals were at a greater risk of ICU admission.
 “Our lab studies have found that non-allergic asthma, or PGA, can suppress immune response to flu and with the immune system compromised, the virus is left unchecked and can replicate more than it does in a healthy individual.
“Our lab studies have found that non-allergic asthma, or PGA, can suppress immune response to flu and with the immune system compromised, the virus is left unchecked and can replicate more than it does in a healthy individual.
“And, since the flu is not so good at proof-reading its genetic code when replicating, it makes a lot of mistakes, and with more replication comes more opportunity for mutations to emerge.”
To conduct the research, the researchers used an asthmatic mouse model with influenza virus.
From there, computer-driven analysis of the virus genome was used to identify mutations that emerged exclusively in the asthmatic group.
UQ’s Dr Kirsty Short said while these tests were preliminary and conducted in animals, the results may reflect a broader phenomenon in humans.
“Our study produced clear findings that fit well with what we know about a suppressed immune response and the emergence of influenza virus variants,” Dr Short said.
“Which is particularly relevant in the context of COVID-19, where it has been suggested that the so-called UK variant arose because of a prolonged infection in an immunocompromised patient.
“For this study, it would be interesting to get access to clinical asthmatic samples to potentially confirm what we’ve found experimentally.
“Our study provides the first evidence that asthma may influence the evolution of the influenza virus, and – transmission permitting – could lead to the emergence of more pathogenic strains into the community.
“It’s therefore really important to remember that host-viral interactions are bidirectional and that host co-morbidities can influence the evolution of influenza virus.”
The research has been published in eLife (DOI: 10.7554/eLife.61803).
Image above left: UQ’s Katina Hulme counting live virus placques.
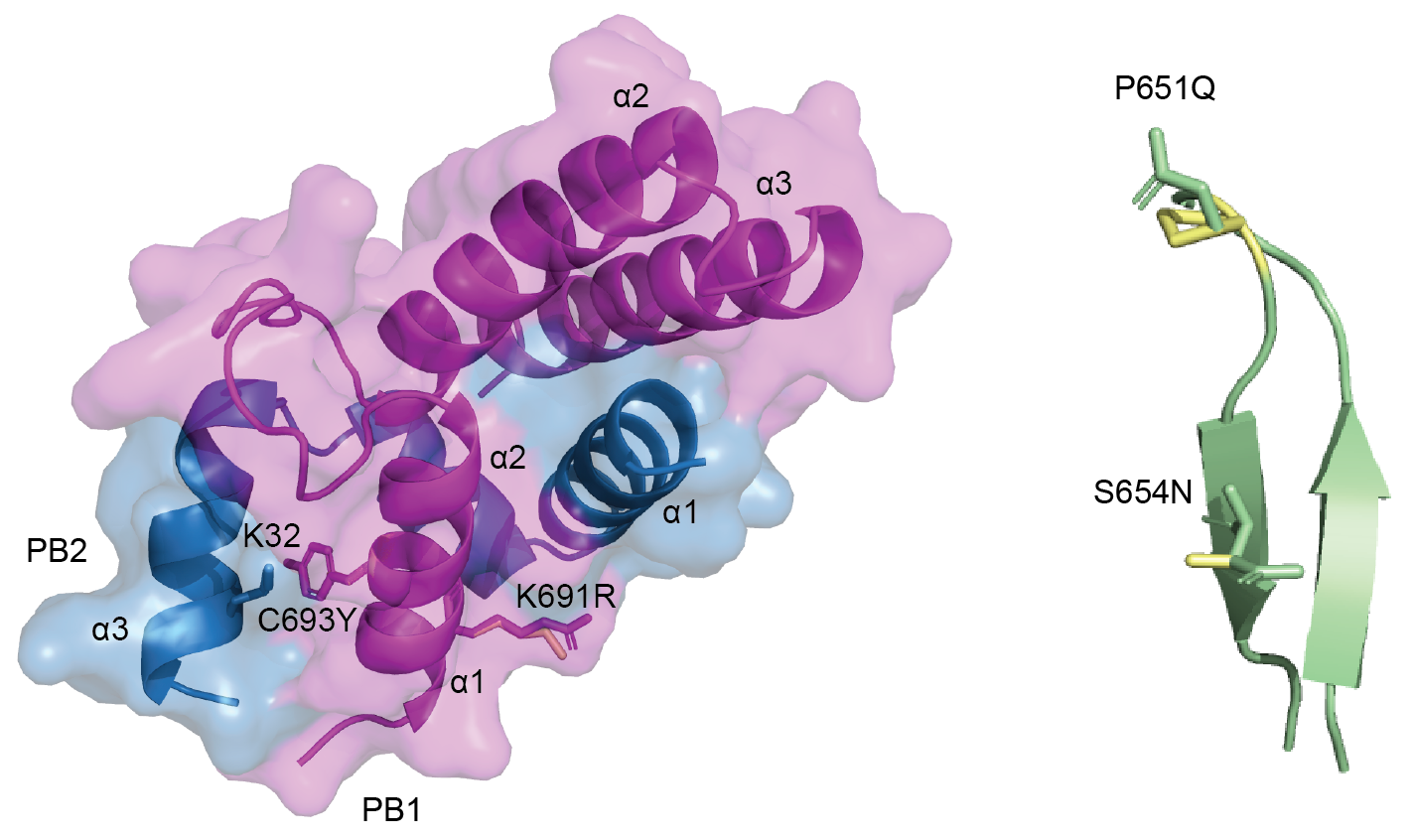
Image above right: computer modelling of influenza gene mutations that were identified in the study.

